GPCA is an algebraic subspace clustering technique based on
polynomial fitting and differentiation. The key idea is to represent
the number of subspaces as the degree of a polynomial and encode the
subspace bases as factors of these polynomials.
Basic GPCA Algorithm
For the sake of simplicity, assume that the data was distributed on a
collection of two hyperplanes, each of one can be represented by the
equation xTbi=0. By multiplying the two equations for the two
subspaces, we obtain
(xTb1)(xTb2)=p(x)=0
where p(x) is a homogeneous polynomial of degree two in the coordinates of
x which is satisfied by all data points, regardless of which
hyperplane they belong to. Therefore this polynomial can be
considered as a global model representing all subspaces.
Fitting of the polynomial
As with any polynomial, p(x) may be expanded linearly in terms of
some coefficients with respect to a basis of polynomials. Therefore,
given enough data points from the subspaces, the coefficients of p(x)
can be linearly estimated from the data by using a non-linear
embedding (the Veronese map). Building upon our previous example, assume that x lives in R3,
say x=[x,y,z]T.
Then, p(x) can be expanded as
p(x)=ax2+by2+cz2+dxy+eyz+fxz.
It is immediate to note that the polynomial is non-linear in the coordinates
x,
y,
z but is linear in the coefficients
a,
b,
c,
d,
e,
f. The latter can be therefore retrived (up to a scale factor) by solving the homogeneous linear system
p(xi)=0, i=1,...,n.
Basis extraction by differentiation
Now, the set of points x such that p(x)=0 can be thought of as a
surface (in our previous example, p(x) is the equation of a conic
in the projective plane P2). Moreover, the gradient of p(x), ∇p(x) must be
orthogonal to this surface. Since the surface is a union of
subspaces, the gradient at a point must be orthogonal to the subspace
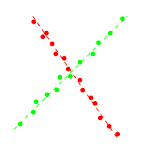
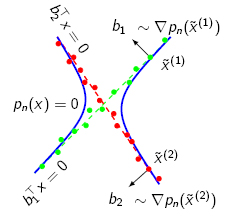
passing through that point. As a result, we may compute the normal
vectors at a point as the gradient of p(x) at that point, as
illustrated in the figure below.
Since all points in the same cluster should have the same (or
similar) normal vectors, we can cluster the data by clustering these
normal vectors. In fact, the angles between these normal vectors
provide use with a similarity measure that can be used together with
spectral clustering.
Extensions of GPCA
Clustering subspaces of different dimensions
In the case of hyperplanes, a single polynomial is enough, but in the case of
subspaces of different dimensions, it is necessary to fit more
polynomials. The gradient of each one of these polynomials at a point
x will give possibly a different normal vector. All these normal
vectors will form a basis for the complement of the subspace passing
through x. The principal angles between these orthogonal subspaces
may now be used to define a similarity measure, and clustering
proceeds as before. More details can be found in the references.
Dealing with noise and outliers
GPCA is an algebraic technique which provide the exact algebraic solution in closed form
when the data is perfect. With a moderate amount of noise, GPCA will
still operate correctly, as the polynomials can be estimated using
least squares. However, in the presence of large amounts of noise and
outliers, the coefficients may be estimated incorrectly. For this
reason, some variations of the main algorithm have been developed.
Some of these employ spectral clustering, voting schemes,
multivariate-trimming and influence functions to deal with non-perfect
data. Please refer to the page from UIUC for more details and code
samples on this topic.
GPCA has been successfully applied in various field of computer
vision. In the following we present a (non-exaustive) list of applications that are based on the ideas of GPCA.
Subspace clustering
[1]
R. Vidal.
Ph.D. Thesis, University of California at Berkeley, 2003.
[2]
R. Vidal, Y. Ma and S. Sastry.
IEEE Conference on Computer Vision and Pattern Recognition, 2003.
[3]
R. Vidal and Y. Ma and J. Piazzi.
IEEE Conference on Computer Vision and Pattern Recognition, 2004.
[4]
K. Huang, Y. Ma, R. Vidal
IEEE Conference on Computer Vision and Pattern Recognition, 2004
[5]
R. Vidal, Y. Ma and S. Sastry.
IEEE Transactions on Pattern Analysis and Machine Intelligence, 2005.
[6]
R. Vidal
International Conference on Machine Learning, 2006
[7]
H. Yi, D. Rajan, L.T. Chia
A ZGPCA Algorithm for Subspace Estimation
IEEE International Conference on Multimedia and Expo, 2007
[8]
D. Zhao, Z. Lin, X. Tang
Contextual Distance for Data Perception
IEEE International Conference on Computer Vision, 2007
[9]
Y. Ma, A. Yang, H. Derksen, R. Fossum
Estimation of Subspace Arrangements with Applications in Modeling and Segmenting Mixed Data,
Preprint, to appear in SIAM Review, 2007
Hilbert series of subspace arrangements
[10]
A. Yang, S. Rao, A. Wagner, Y. Ma, R. Fossum
Hilbert Functions and Applications to the Estimation of Subspace Arrangements
IEEE International Conference on Computer Vision, 2005
[11]
H. Derksen
Hilbert series of subspace arrangements
Journal of Pure and Applied Algebra, 2007
Motion segmentation
[12]
R. Vidal, S. Sastry
IEEE Workshop on Vision and Motion Computing, 2002
[13]
R. Vidal, S. Sastry
IEEE Conference on Computer Vision and Pattern Recognition, 2003
[14]
R. Vidal, R. Hartley
IEEE Conference on Computer Vision and Pattern Recognition, 2004
[15]
F. Shi, J. Wang, J. Zhang, Y. Liu
Motion Segmentation of Multiple Translating Objects Using Line Correspondences
IEEE Computer Society Conference on Computer Vision and Pattern Recognition, 2005
[16]
J. Yan, M. Pollefeys
A Factorization-Based Approach for Articulated Non-rigid Shape, Motion and Kinematic Chain Recovery
IEEE Conference on Computer Vision and Pattern Recognition, 2005
[17]
R. Vidal, Y. Ma
Journal of Mathematical Imaging and Vision, 2006
[18]
R. Vidal, Y. Ma, S. Soatto, S. Sastry
International Journal of Computer Vision, 2006
[19]
R. Vidal
Imaging Beyond the Pinhole Camera, Springer, 2006
[20]
R. Tron, R. Vidal
IEEE International Conference on Computer Vision and Pattern Recognition, 2007
[21]
R. Vidal
IEEE Transactions on Pattern Analysis and Machine Intelligence, 2007
[22]
J. Zhang, F. Shi, J. Wang, Y. Liu
3D Motion Segmentation from Straight-Line Optical Flow
Multimedia Content Analysis and Mining, Springer, 2007
Hybrid Systems
[23]
R. Vidal, S. Soatto, Y. Ma, S. Sastry
IEEE Conference on Decision and Control, 2003
[24]
R. Vidal
American Control Conference, 2004.
[25]
R. Vidal, B.D.O. Anderson
IEEE Conference on Decision and Control, 2004
[26]
Y. Hashambhoy and R. Vidal
IEEE Conference on Decision and Control, 2005
[27]
R. Vidal, S. Soatto, A. Chiuso
European Control Conference, 2007
[28]
R. Vidal
Recursive Identification of Switched ARX Systems
Automatica, 2007
Dynamic Textures
[29]
R. Vidal, A. Ravichandran
IEEE Conference on Computer Vision and Pattern Recognition, 2005
[30]
A. Ghoreyshi, R. Vidal
Dynamical Vision Workshop in the European Conf. on Computer Vision, 2006
[31]
L. Cooper, J. Liu, K. Huang
Spatial segmentation of temporal texture using mixture linear models
Dynamical Vision, Springer, 2007
Image segmentation and compression
[32]
J. Ye, R. Janardan, Q. Li
GPCA: an efficient dimension reduction scheme for image compression and retrieval
ACM SIGKDD international conference on Knowledge discovery and data mining, 2004
[33]
W. Hong, J. Wright, K. Huang, Y. Ma
A Multi-Scale Hybrid Linear Model for Lossy Image Representation
IEEE International Conference on Computer Vision, 2005
[34]
R. Samur, V. Zagorodnov
Segmenting Small Regions in the Presence of Noise
IEEE International Conference on Image Processing, 2005
[35]
Y. Hu, D. Rajan, L.T. Chia
Scale adaptive visual attention detection by subspace analysis
International Conference on Multimedia, 2007
Biomedical imaging
[36]
A. Ravichandran, R. Vidal, H. Halperin
Segmenting a Beating Heart Using Polysegment and Spatial GPCA
IEEE International Symposium on Biomedical Imaging: Nano to Macro, 2006
Vision-Based Formation Control
[37]
R. Vidal, O. Shakernia and S. Sastry
Following the Flock: Distributed Formation Control with Omnidirectional Vision-Based Motion Segmentation and Visual Servoing.
IEEE Robotics and Automation Magazine, 2004
Please refer to the specific webpages on this same website for more
details on this topic.